Data collection and pre-processing
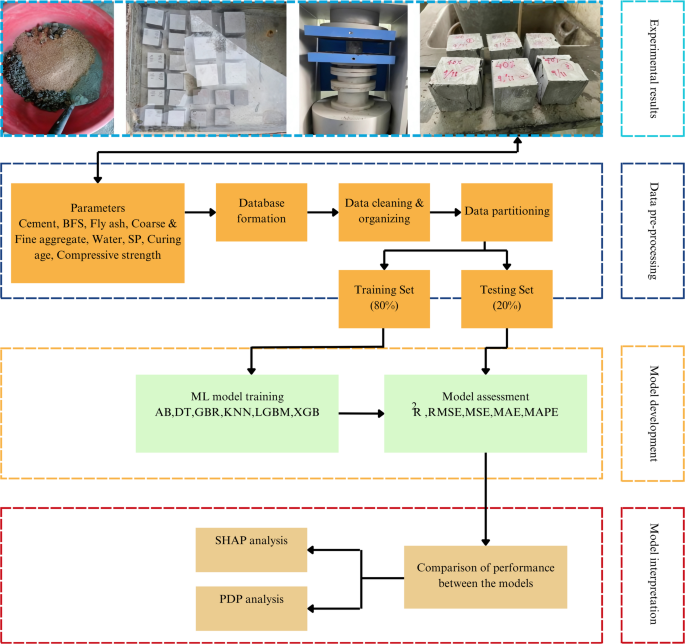
In the present study, a thorough examination of existing research on BFS incorporated concrete was conducted to construct the algorithmic predictors of CS. A comprehensive database of 675 records has been curated through an in-depth analysis of published experimental studies and ML repositories, which contain verified experimental datasets on BFS incorporated concrete77,78,79,80,81,82,83,84,85,86,87,88,89. The data collection involved systematically reviewing prior research that reported CS and corresponding mix composition parameters used in this research. Each dataset was extracted, cross-checked, and standardized to maintain consistency in units, mix proportions, and test conditions across different studies. Then, they were compiled into an Excel sheet, where they were systematically organized, categorized, and subjected to statistical analysis. An overview of the methodology is shown in Fig. 1.
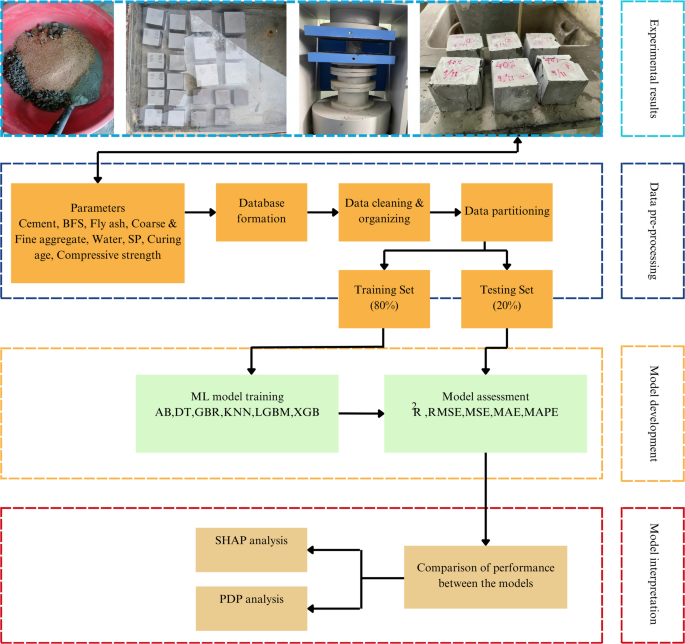
Flowchart of the study methodology.
For the ML simulations, the compiled dataset was partitioned using a random split into train/test sets. A total of 80% of the data was designated for the training set, utilized to develop the model, and validated through cross-checking for optimal hyperparameter adjustment. The data cross-checking process involved two authors independently re-entering 10% of the rows sampled at random and comparing them against the compiled sheet (acceptance threshold: ≤0.5% relative deviation per field). Discrepancies triggered a full re‑check of the source paper. A third author validated all unit conversions. The leftover 20% of the compiled data was reserved as the testing set, employed to assess the effectiveness and accuracy of the developed models. The laboratory data referenced from the literature were collected in a randomized manner to ensure an unbiased representation. Although the dataset encompassed numerous variables, only those parameters with a significant influence on the mechanical properties were chosen and subsequently processed. While many of these experimental studies also assess other concrete properties, this research focused solely on CS related data points, which were selectively gathered in sufficient quantity to support model development.
Eight input variables were used to develop the models, including the amount of cement, BFS, fly ash, coarse aggregate, fine aggregate, water, SP, and curing period, while CS was the output variable. To ensure consistency and minimize potential errors introduced during the simulation process, all data values were meticulously transformed and standardized to a uniform scale ranging from 0 to 1. This normalization step was integral to the model construction, facilitating enhanced computational efficiency and improved accuracy in the simulation outcomes. Data manipulation and preprocessing employed data analysis libraries of Pandas v2.2 and NumPy v1.26, with statistical computations facilitated by SciPy v1.12.
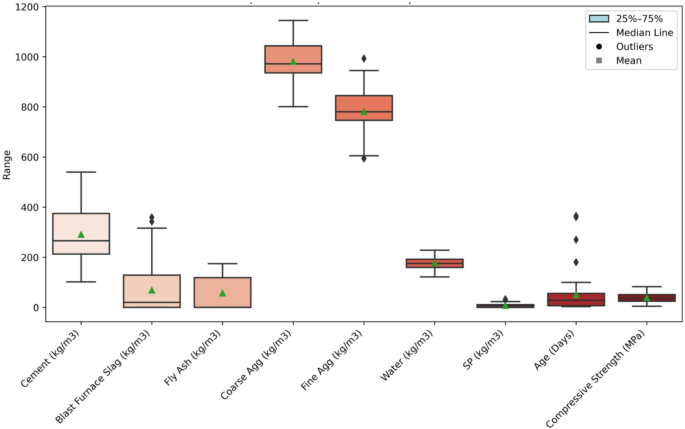
The additional data processing steps encompassed data cleaning, handling missing values, encoding categorical data and treating outliers. The statistical summary of all variables is presented in Table 2. Outliers were detected using the interquartile range (IQR) method, where values below \(\:Q1-1.5\times\:IQR\) or above \(\:Q3+1.5\times\:IQR\) were identified as extremes. Rather than deleting these values, they were winsorized to the nearest valid limit to maintain the full dataset size. Figure 2 illustrates the variable distributions and the IQR-based outlier detection process applied to each parameter. Together, these steps ensured a validated and balanced dataset suitable for reliable model training.
Boxplots of input and output variables.
Data representation
Data representation is an essential tool for understanding complex datasets. It allows for the effective communication of patterns, relationships, and trends that may not be obvious through experimental data alone90. In this study, we employ histograms with overlaid density plots and a correlation heatmap to explore the frequency profiles of input and output features and their interrelationships. A comprehensive summary presented in Table 2 enumerates the key distribution statistics for each input and output variable in the dataset, including mean, standard deviation, quartiles, and observed ranges.
In addition, the boxplot visualization referred to as Fig. 2 concisely displays the central tendency, dispersion, and outlier distribution for all variables. Cement content has a mean of 290.74 kg/m3, with values spreading from 102 kg/m3 to 540 kg/m3, indicating a wide variety of mix designs. Figure 2 highlights a concentration of values around 266 kg/m3, with some extreme outliers. BFS shows similar variability, with a mean of 67.99 kg/m3 ranging from 0 to 359.4 kg/m3 with frequent outliers. Water content ranges from 121.8 kg/m3 to 228 kg/m3, while SP is typically used in small amounts, showing less variability. Coarse and fine aggregates show extensive interquartile ranges, with means of 981.21 kg/m3 and 780.93 kg/m3, respectively, are more consistent across the samples.
CS ranges from 4.57 MPa to 82.6 MPa with a median of 37.4 MPa and several high‑strength outliers, features that the whiskers and mean markers in Fig. 2 display clearly. This wide range can be linked to the variations in input materials. Higher cement content and the presence of BFS seem to positively affect the CS. Concrete mixes with higher cement content and optimal amounts of slag generally show stronger CS. The consistent use of water, SP and aggregates further influences the overall strength, showing that the mix design directly impacts the concrete’s performance.
Figure 3 presents a series of histograms with density plots for each input variable. These visualizations highlight key aspects of the data distribution and provide a clearer understanding of how each variable behaves. Cement shows a relatively normal distribution, with values mostly centered around 200 kg/m3 to 400 kg/m3. This suggests that cement content is fairly consistent across the dataset, although a few higher values indicate mixes that use a larger proportion of cement.
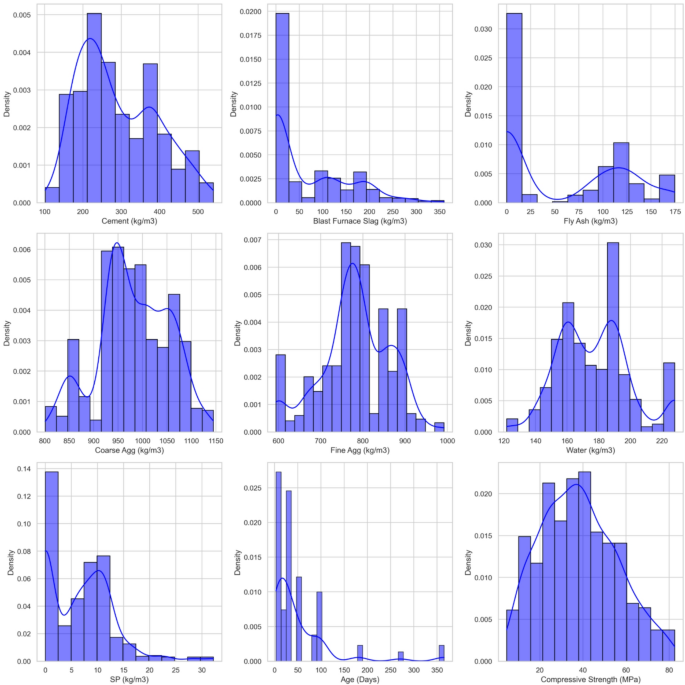
Histograms and density plots of dataset variables.
BFS results in a more prominent distribution. The histogram reveals a concentration of values between 0 and 150 kg/m3, followed by a gradual decrease as slag content increases. This indicates that lower amounts of BFS are more commonly used in the mixes, although there are some concrete designs containing higher levels of slag. The density plot clarifies that while most mixes contain low slag, a smaller subset uses high quantities. This difference may impact cost and material properties (e.g., strength, durability). Fly Ash follows a similar behavior to slag but shows a slightly more skewed distribution, with values ranging from 0 to 174.7 kg/m3. Water content is near symmetric around 175 to 190 kg/m3. Its slight upward skew indicates a common trend toward higher-water mixes. SP usage, with a strong right-skewed distribution, is consistent with dosage-on-demand practice. Curing age is multi-modal with peaks at standard curing times (notably 28 days) and secondary modes at extended ages. Finally, CS is unimodal with a slight positive skew, indicating more mid-strength than very high-strength mixes.
Figure 4, a correlation heatmap, provides visibility into the interactions between these variables, illustrating both positive and negative correlations. The heatmap illustrates that cement shows a positive correlation of about 0.48 with CS, confirming that higher cement content generally produces stronger concrete. BFS exhibits a weak positive correlation of around 0.19 with CS, suggesting that slag provides a modest improvement in strength. In contrast, fly ash, coarse aggregate, fine aggregate and water show slight to moderate negative correlations with CS. SP and curing age demonstrate strong positive correlations of 0.46 and 0.30, respectively, emphasizing their substantial role in strength enhancement.
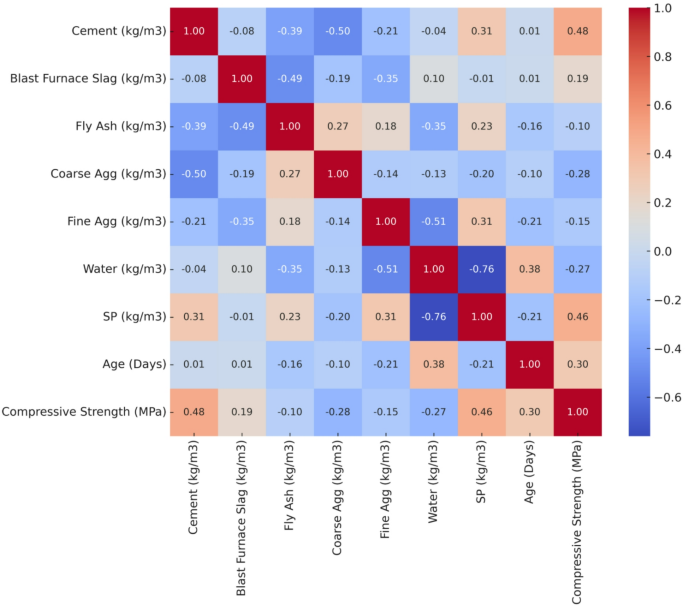
Correlation heatmap of concrete mix parameters and CS.
Model development
This research investigated a range of algorithms to select the best-performing model for concrete incorporating BFS. In this study, classical regression models such as linear and polynomial regression were not adopted because they rely on rigid functional assumptions that struggle to represent the complex, nonlinear interactions inherent in BFS-incorporated concrete. Relationships among binder, aggregate, SCMs, and curing kinetics with CS are highly non-linear and exhibit interdependent effects that cannot be adequately captured through linear combinations of parameters or simple transformations. In contrast, ensemble tree-based methods and gradient boosting frameworks can model nonlinear feature interactions without prespecifying functional forms, which makes them more appropriate for accurately learning composition-strength relationships. Therefore, classical regression approaches were excluded at the model selection stage to ensure methodological alignment with the underlying material behavior.
The evaluated algorithms included AdaBoost, DT, GBR, XGBoost, KNN and LightGBM. Each model was evaluated using a random 80:20 train-test split and a statistical metrics approach to ensure the generation of the most reliable results. The analysis incorporated eight input variables and a single output variable, as detailed in the preceding sections. All computational analyses were performed on a Windows 11 64-bit system using Python 3.11. Model development and validation were conducted using scikit-learn version 1.7, while XGBoost version 2.0 and LightGBM version 4.3 enabled advanced ensemble modeling. Visualization outputs were generated using Matplotlib v3.8, Seaborn v0.13, and SHAP v0.45.
Adaptive boosting (AdaBoost/ADB)
AdaBoost is an ensemble algorithm that facilitates predictive accuracy by concentrating on tough examples91. For regression, AdaBoost combines multiple weak learners \(h_ – 1 \left( x \right)\) by reweighting samples according to their relative absolute error92. For iteration \(t\) with sample weights \(w_ s \right^{\left( t \right)}\),
$$e_{i} = \frac{{\left| {y_{i} – h_{t} \left( {x_{i} } \right)} \right|}}{{R_{t} }}$$
$$R_{t} = \max \left| {y_{j} – h_{t} \left( x \right)} \right|$$
The weighted error and stage parameter are,
$$\varepsilon_{t} = \mathop \sum \limits_{i = 1}^{N} w_{{\dot{i}}}^{\left( t \right)} e_{i}$$
$$\beta_{t} = \frac{{\varepsilon_{t} }}{{1 – \varepsilon_{t} }}$$
Weights are updated and normalized as,
$$w_{i}^{{\left( {t + 1} \right)}} = w_{i} \beta_{t}^{{1 – e_{i} }}$$
$$\mathop \sum \limits_{i} w_{i}^{{\left( {t + 1} \right)}} = 1$$
Each learner receives weight, \(\alpha_{t} = \ln \left( {1{ / }\beta_{t} } \right)\). The final ensemble prediction is the weighted median of all weak learners,
$$\hat{y}\left( x \right) = median~(~h_{1} \left( x \right),….,h_{T} \left( x \right);\alpha _{1} ,….,\alpha _{T} ).$$
Decision tree (DT)
DT is widely utilized for classification and regression tasks. They work by repeatedly splitting the data based on features, attempting to reduce variance at each node93. DT for regression splits the data to minimize the sum of squared errors (SSE) at each node94. For a node \(P\) containing samples \(\left\{ {\left( {x_{i} ,y_{i} } \right)} \right\}_{iP}\) are,
$$I\left( P \right) = \mathop \sum \limits_{i \in P} \left( {y_{i} – \overline{y}_{P} } \right)^{2}$$
$$\overline{y}_{P} = \frac{1}{{n_{P} }}\mathop \sum \limits_{i \in P} y_{i}$$
A candidate split \(s\) on a feature produces child nodes \(L,R\). The reduction (gain) in impurity is,
$$\Delta I\left( s \right) = I\left( P \right) – \left[ {I\left( L \right) + I\left( R \right)} \right]$$
The algorithm chooses the split that maximizes \(\Delta I\left( s \right)\) and continues recursively until a stopping criterion (e.g., maximum depth, minimum samples per leaf) is satisfied. The final leaf prediction is the mean of targets in that leaf,
$$\hat{y}\left( x \right) = \overline{y}_{leaf\left( x \right)}$$
Gradient boosted regression (GBR)
GBR is a highly effective ML algorithm that combines the principles of boosting and regression to deliver accurate predictive models93. At the start, it creates a simple model \(F_{0}\) (often a constant) that minimizes the chosen loss function \(L\). In each round \(m\), it calculates the “pseudo-residuals” \(r_{im}\) for each data point,
$$r_{im} = – \left[ {\frac{{\partial L\left( {y_{i} ,F\left( {x_{i} } \right)} \right)}}{{\partial F\left( {x_{i} } \right)}}} \right]_{{F = F_{m – 1} }}$$
where \(F_{m – 1}\) is the model from the previous step, and \(y_{i}\) is the true target. It then trains a weak regressor \(h_{m}\) (commonly a shallow tree) on these residuals. Next, the algorithm finds the multiplier \(\gamma_{m}\) that best fits the new weak regressor to reduce the loss. Finally, it updates the model:
$$F_{m} \left( x \right) = F_{m – 1} \left( x \right) + \nu \gamma_{m} h_{m} \left( x \right)$$
where \(\nu\) is the learning rate that controls how quickly the model learns. Repeating this process for \(M\) iterations yields a final model that sums all weak regressors. By continually adjusting for past errors, GBR steadily boosts predictive accuracy46.
k-Nearest neighbors (KNN)
KNN is based on the principle of proximity, where the outcome for a given data point is determined by the modal class or mean value of its -nearest neighbors in the predictor space. It is a model-free method that makes no specific assumptions about the underlying distribution95. KNN predicts CS for a query point \(x_{q}\) as the average strength of its \(k\) nearest neighbors,
$$\hat{y}\left( {x_{q} } \right) = \frac{1}{k}\mathop \sum \limits_{{i \in N_{k} \left( {x_{q} } \right)}} y_{i}$$
where distance is typically Euclidean,
$${\text{d}}\left( {x_{i} ,x_{q} } \right) = \sqrt {\mathop \sum \limits_{j = 1}^{p} \left( {x_{ij} – x_{qj} } \right)^{2} }$$
Here, \(x_{{\dot{i}}}\) is a training instance described by \(p\) features. The \(k\) closest samples (i.e., those with the smallest distances) form the “neighborhood.” In classification, KNN typically chooses the highest frequency data among these \(k\) neighbors; for regression, it often takes the mean of their target values. The parameter \(k\) significantly influences performance, small values may overfit to local noise, while large values risk over-smoothing predictions. Because KNN postpones its core calculations until it needs to predict a query point96.
Light gradient boosting machine (LightGBM/LGB)
LightGBM is a powerful and sample-efficient algorithm for gradient boosting tasks. It builds decision trees by focusing on the leaf that achieves the greatest improvement in the loss function. Instead of adding a tree layer by layer, LightGBM performs leaf-wise growth, which enables it to capture more complex patterns in the data97. To update the model, LightGBM adds a new tree \(h_{m} \left( x \right)\) to the previous model \(F_{m – 1} \left( x \right)\) with a learning rate \(\nu\):
$$F_{m} \left( x \right) = F_{m – 1} \left( x \right) + \nu \cdot h_{m} \left( x \right)$$
Defining first- and second-order gradients of the loss,
$$g_{i} = \frac{{\partial L\left( {y_{i} ,\hat{y}_{1}{\prime} } \right)}}{{\partial \hat{y}_{i} }}$$
$$h_{i} = \frac{{\partial^{2} L\left( {y_{i} ,\hat{y}_{i} } \right)}}{{\partial \hat{y}_{i}^{2} }}$$
$$\hat{y}_{i} = F_{m – 1} \left( {x_{i} } \right)$$
For any node, the gradient and Hessian sums are defined as,
$$G = \Sigma _{{i\varepsilon node}} g_{i} \;and\,H = \Sigma _{{i\varepsilon node}} h_{i}$$
The gain for a split into left (L) and right (R) children is,
$$Gain = \frac{1}{2}\left( {\frac{{G_{L}^{2} }}{{H_{L} + \lambda }} + \frac{{G_{R}^{2} }}{{H_{R} + \lambda }} – \frac{{\left( {G_{L} + G_{R} } \right)^{2} }}{{H_{L} + H_{R} + \lambda }}} \right) – \gamma$$
where \(\lambda\) represents the \(L_{2}\) regularization term for leaf weights and \(\gamma\) is a complexity penalty that discourages overfitting. LightGBM continues leaf-wise growth until the stopping criteria are met.
Extreme gradient boosting (XGBoost/XGB)
XGBoost assembles a series of decision trees, where each iteration corrects the residual errors of previous ones. XGBoost applies a second-order Taylor expansion to optimize the objective function, which allows for faster convergence and more accurate models98.
At boosting stage \(t\), adding a new tree \(f_{t} \left( x \right)\) to the current prediction \(\hat{y}_{i}^{{\left( {t – 1} \right)}}\)gives,
$$Obj^{\left( t \right)} = \mathop \sum \limits_{i = 1}^{N} L\left( {y_{i} ,\hat{y}_{i}^{{\left( {t – 1} \right)}} + f_{t} \left( {x_{i} } \right)} \right) + \Omega \left( {f_{t} } \right)$$
with tree complexity,
$$\Omega \left( {f_{t} } \right) = \gamma T + \frac{1}{2}\lambda \mathop \sum \limits_{j = 1}^{T} w_{j}^{2}$$
where \(T\) is the number of leaves and \(w_{j}\) is the weight of leaf \(j\). Using the second-order expansion around \(\hat{y}_{i}^{{\left( {t – 1} \right)}}\) with,
$$g_{i} = \left. {\frac{{\partial L\left( {y_{i} ,\hat{y}} \right)}}{{\partial \hat{y}}}} \right|_{{\hat{y} = \hat{y}_{i}^{{\left( {t – 1} \right)}} }} , h_{i} = \left. {\frac{{\partial^{2} L\left( {y_{i} ,\hat{y}} \right)}}{{\partial \hat{y}^{2} }}} \right|_{{\hat{y} = \hat{y}_{i}^{{\left( {t – 1} \right)}} }}$$
Collecting samples of leaf \(j\) in the index set \(I_{j}\), the stage objective (up to constants) becomes,
$$O\tilde{b}j^{\left( t \right)} = \mathop \sum \limits_{j = 1}^{T} \left[ {\mathop \sum \limits_{{i \in I_{j} }} \left( {g_{i} w_{j} + \frac{1}{2}h_{i} \omega_{j}^{2} } \right)} \right] + \gamma T + \frac{1}{2}\lambda \mathop \sum \limits_{j = 1}^{T} w_{j}^{2}$$
The optimal leaf weight and the corresponding gain are,
$$w_{j}^{*} = \frac{{\Sigma_{{i \in I_{j} }} g_{i} }}{{\Sigma_{{i \in I_{j} }} h_{i} + \lambda }}$$
$$Gain = \frac{1}{2}\mathop \sum \limits_{j = 1}^{T} \frac{{\left( {\Sigma_{{i \in I_{j} }} g_{i} } \right)^{2} }}{{\Sigma_{{i \in I_{j} }} h_{i} + \lambda }} – \gamma T$$
After fitting \(f_{t}\), the model is updated with learning rate \(\nu\),
$$F_{t} \left( x \right) = F_{t – 1} \left( x \right) + \nu f_{t} \left( x \right)$$
Hyperparameter tuning
We tuned hyperparameters to improve model accuracy and prevent overfitting. For the DT model, Fig. 5 illustrates RMSE versus tree depth that tested different tree depths. As depth increased, training RMSE dropped sharply, showing the model fit the training data better. However, after a certain depth, the testing RMSE stopped decreasing and even started to rise slightly. This indicated that the model was initializing to track the training data rather than learning general patterns. We selected the tree depth where the testing RMSE became stable. This approach gave us a model that was accurate on unseen data without being too complex.
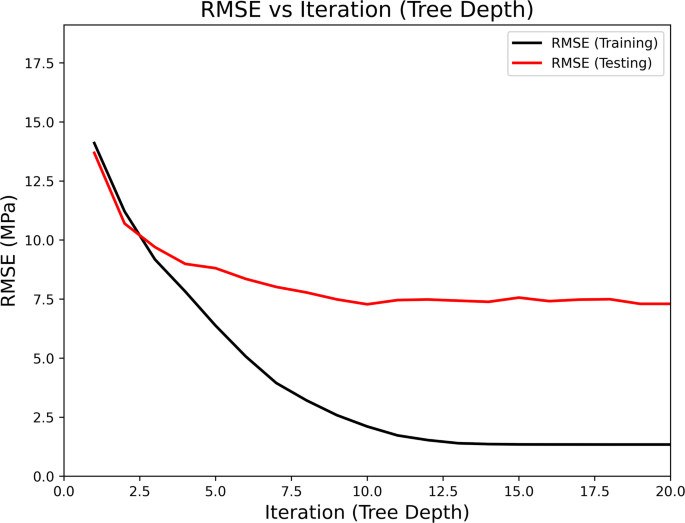
Variation of RMSE with tree depth in the DT model.
Similarly, for the KNN regressor, Fig. 6 displays mean error versus K value. At low k values, the model had low training error but higher validation error, suggesting overfitting. As we increased k, the training error rose, but the validation error initially decreased, reaching its lowest point at k = 3. Beyond this, the validation error started to climb, indicating underfitting as the model became too simple. The U-shaped curve of validation error across k values helped us clearly identify the optimal k. We chose the k value that gave the lowest validation error, ensuring the model could generalize well. Throughout the tuning process, we relied on trends in both training and validation errors. This helped us avoid both underfitting and overfitting. The final hyperparameter choices for each model were based on the points where validation performance was best.
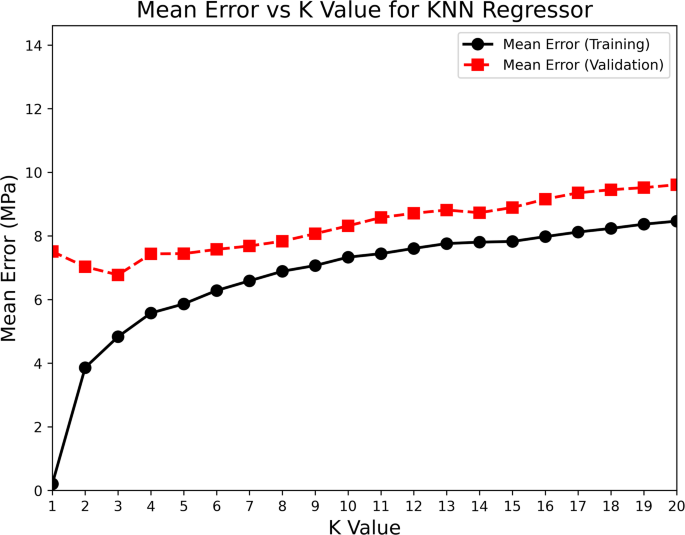
Mean prediction with different values of \(\:K\) in the KNN algorithm.
Model accuracy assessment
In this study, a comprehensive set of performance metrics was used to assess the predictive accuracy of the ML models applied to concrete strength prediction. The chosen metrics—MAE, MAPE, MSE, RMSE, R2 provide a thorough evaluation of model accuracy, reliability, and generalizability across various datasets.
MAE calculates the average absolute differences between the predicted and actual values. This straightforward metric offers a clear interpretation of how far off the model’s predictions are on average, without giving excessive weight to outliers99. MAPE provides a measure of prediction accuracy that is easy to interpret across datasets with different scales100. MSE reflects the average squared difference between actual and predicted values. It gives more importance to larger errors than MAE and provides insight into prediction errors101. RMSE, a derived metric from MSE, further emphasizes larger discrepancies by taking the square root. A lower RMSE indicates better model accuracy by reducing the deviation from actual values102. Finally, R2 measures how well the model explains the variance in the target variable. A higher R-squared value demonstrates better model fit, with values closer to 1, meaning a stronger predictive relationship103.
These metrics are instrumental in comparing model performance. In this study, they are used to evaluate how well each ML model fits the data and how accurately it predicts CS in BFS-incorporated concrete. A combination of these metrics covers multiple model performance aspects. These include error magnitude, accuracy, and the proportion of variance. The equations for each metric are as follows:
$$MAE = \frac{1}{N}\mathop \sum \limits_{i = 1}^{N} \left| {y_{i} – \hat{y}_{i} } \right|$$
$$MAPE = \frac{100}{N}\mathop \sum \limits_{i = 1}^{N} \left| {\frac{{y_{i} – \hat{y}_{i} }}{{y_{i} }}} \right|$$
$$MSE = \frac{1}{N}\mathop \sum \limits_{i = 1}^{N} (y_{i} – \hat{y}_{i} )^{2}$$
$$RMSE = \sqrt {\frac{1}{N}\mathop \sum \limits_{i = 1}^{N} (y_{i} – \hat{y}_{i} )^{2} }$$
$$R^{2} = 1 – \frac{{\mathop \sum \nolimits_{i = 1}^{N} \left( {y_{i} – \hat{y}_{i} } \right)^{2} }}{{\mathop \sum \nolimits_{i = 1}^{N} \left( {y_{i} – \overline{y}} \right)^{2} }}$$
where \(y_{i}\) is the actual value, \(\hat{y}_{i}\) is the predicted value, and \(\overline{y}\) is the mean of the observed values.
Model interpretation with SHAP
SHAP values are used to analyze ML models by linking each feature a contribution to the prediction104. For a model \(f\left( x \right)\) with \(M\) features, the prediction can be decomposed as,
$$f\left( x \right) = \phi_{0} + \mathop \sum \limits_{j = 1}^{M} \phi_{j} \left( x \right)$$
where \(\phi_{0} = {\mathbb{E}}\left[ {f\left( X \right)} \right]\) is the baseline (mean prediction over the dataset), and \(\phi_{j} \left( x \right)\) is the contribution of the feature \(j\) for sample \(x\). The Shapley value for each feature is computed as,
$$\phi_{j} \left( x \right) = \mathop \sum \limits_{{s \subseteq F{ \setminus }\left( j \right)}} \frac{{\left| s \right|!\left( {M – \left| s \right| – 1} \right)!}}{M!}\left[ {f_{{s \cup \left\{ j \right\}}} \left( x \right) – f_{s} \left( x \right)} \right]$$
where \(s\) represents a subset of all features, excluding \(j\), and \(f_{s} \left( x \right)\) is the expected model output when only the features in \(s\) are fixed to their observed values.
For overall global importance, the mean absolute SHAP value is calculated as,
$$GlobalImportance\left( j \right) = \frac{1}{N}\mathop \sum \limits_{i = 1}^{N} \left| {\phi_{j} \left( {x^{\left( i \right)} } \right)} \right|$$
The model’s overall performance mainly depends on features with higher average SHAP values. This allows for the identification of the key factors behind the model’s predictions105. SHAP values explain the specific contribution of each feature to an individual prediction by local interpretation. Positive SHAP values indicate that the feature increases the predicted outcome, whereas negative values suggest a reducing effect on the prediction106. This interpretation helps in understanding the model’s decision-making process and enhances its usability in practical applications.
Model interpretation with PDP
PDP displays the effect of a single feature on model predictions while keeping other features constant. PDP offer a clear view of how a feature influences the output across its entire range by visualizing the relationship107. For a feature \(x_{j}\), the partial dependence is measured by averaging the predictions \(f\left( x \right)\) over the dataset, while fixing the values of all other features through this formula:
$$PDP\left( {x_{j} } \right) = \frac{1}{N}\mathop \sum \limits_{i = 1}^{N} f\left( {x_{1} ,x_{2} , \ldots ,x_{j – 1} ,x_{j} ,x_{j + 1} , \ldots ,x_{p} } \right)$$
This plot illustrates how the prediction changes as the value of \(x_{j}\)varies. PDPs are especially effective for examining non-linear relationships between features and the target variable108. In this study, PDPs provide a clear and visual way to explore the role of each feature in guiding model predictions. They help clarify complex relationships and improve the model’s transparency and interpretability.
link